- in Blog Articles by Sam Driver
Know when to fold ’em
With my background in genetics, protein biochemistry was never my favorite class. There was always a friendly rivalry between genetics and biochemistry, and it didn’t help that the biochemists always had prettier molecules than we did in our DNA and RNA. So leave it to a team of biochemists to rub it in with the introduction of Fold-It (see Figure below). Sour grapes aside, I downloaded and played the game. It’s fun – the game aspects, ease of use, nice visuals, rapid play and positive feedback and rewards engaged me. The protein structures are rendered in bright colors and the sum total is enough to get you to forget you are doing (yawn) chemistry.
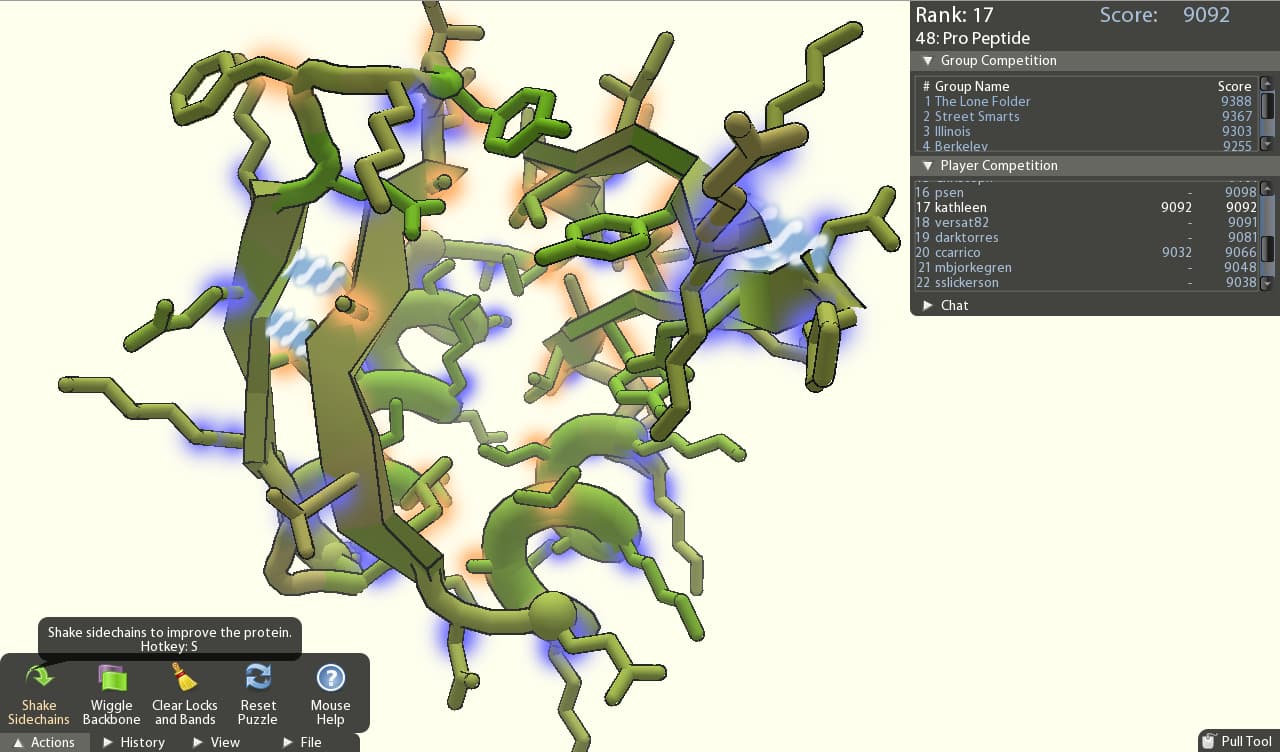
You might expect that the scientific community is using a game to try to educate students and entice them into the field, but what makes this story so interesting is that is not actually the stated goal of the game developers at the University of Washington. In fact, the main idea behind the game is to reach out into the larger world and access the imaginations, creativity, and brain power of more people than the scientific community would normally have access to, in order to solve complex problems. Many more people. This is important because:
- There aren’t enough scientists or powerful enough computers to figure it all out. If you assume that any given protein folding problem is probably only looked at in earnest by dozens or even a couple of hundred people worldwide, you place a lot of pressure on these individuals to visualize complex folding patterns. Computers can’t do it this visualization as well as people can, so the game is an effort to bring new sets of eyes and the brains behind them to solving the puzzles.
- Solutions may come from unexpected sources. Extending the workforce from maybe a couple hundred scientists to perhaps tens of thousands of interested individuals will bring the time required to find a solution way down, tapping into visualization geniuses who may never have studied chemistry in school. This is an exceedingly cool solution to a limited-resource problem, which is a central inhibitor to nearly all scientific research. It illustrates the potential of using a global, multi-user 3D environment to increase the pace of drug discovery, and it creates a fun game for people who may not have any interest in biochemistry to really help the cause, if only by accident.
The big question looming in my mind centers around the intellectual property that may develop out of this game. An often joked about lawsuit, Antonio Hernandez vs. Internet Gaming Entertainment Ltd., in the massively multiplayer online role-playing game (MMORPG) sector, could point at the center of the issue: do players have any IP rights to discoveries made in the game environment? This may not be the particular legal case that decides the matter, but when you are talking about potentially discovering something that is turned into a diagnostic tool or pharmaceutical, be assured there will be legal wrangling. But threats of lawsuits should never hamper innovation. So I’m heading back to my desk to fold some more proteins. Come join me!
